|
All living cells of plants and animals have mitochondria, organelles that convert carbohydrates into the useful form of energy that drives synthesis of metabolites in cells. Mitochondria are believed to be descendants of bacteria that became symbiotic with cells in the early evolution of most living forms. They retained their own DNA, are transferred to the next generation only in eggs cell and not sperm. They replicate within cells but the host cells have some control on the rate of replication. Energy conversion in mitochondria occurs on their folded membranes in a series of chemical reactions. Regions of the plant undergoing rapid cell duplication have more mitochondria. This includes the tassel cells of a corn plant. The pollen mother cells in that region undergo meiosis and duplication, driven partly by the energy conversion by concentration of mitochondria in those mother cells.
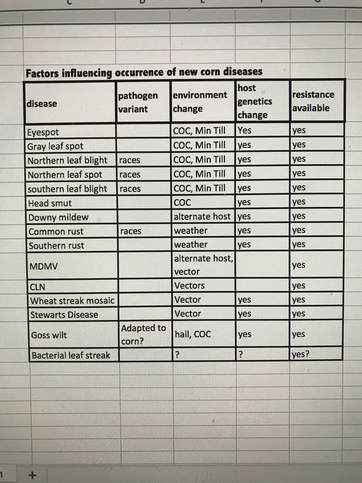
A small defect in mitochondrial DNA of an inbred caused a defective membrane product in those mitochondria resulting in incomplete development of pollen. This was found in a corn breeding program in Texas. As the inheritance of this condition was known to be only transmitted independent of nuclear DNA, it was called Texas male sterile cytoplasm. It became a useful tool to corn hybrid seed production because it was easily transferred in breeding programs to the female parent of a hybrid, and thus avoiding manual removal of tassels in seed production fields. Use of T male-sterile cytoplasm became common in the worldwide corn in the 1960’s. It was noted in the Philippines in 1961, that a fungal pathogen, then known as Helminthosporium maydis, was especially aggressive on several hybrids with T cytoplasm. Despite a few scattered reports elsewhere it was not until 1969 that the connection between increased occurrence of this disease and T cytoplasm became alarming. Majority of seed produced for 1970 corn season had T cytoplasm, the main exceptions being new hybrids in which the conversion to sterility of the female parents was incomplete. Although the pathogen was normally found in the southern half of the corn belt, and adequately controlled by products of nuclear DNA genes, this disease was found highly destructive in northern corn belt areas as well. A race of the fungus (now named Bipolaris maydis and by its sexual stage Cochliobolus heterostrophus) called race T, produces a toxin that causes death to cells with mitochondria having the DNA with the defect associated with T male sterility. All cells of the corn plant with these defective mitochondria were vulnerable to the fungus. This included the cells in developing seed resulting in diseased stored grain as well as overwintering leaves and stalks. Normal resistance mechanisms to the pathogen were ineffective because the toxin destroyed these defective mitochondria. As the relationship with T cytoplasm was realized, seed companies worked to change, and within a few years, the disease subsided back to its normal distribution. It was a new learning experience of interaction of corn and pathogen biology. During my short time (uh, 46 years) of looking at corn diseases in the U.S.A. and some time elsewhere, it's interesting, at least to me, that every few years a new corn disease problem is observed. With each, it takes a few years for the private and public researchers to understand the dynamics that led to the new problem and the solutions. Analysis always comes down to some aspect of the pathogen, host and environment triangle as taught in the beginning of plant pathology courses. Pathogens have their genetics, with variants, survival pressures dependent on feeding on dead and/or alive plant tissue and preference for favorable environment. Environments for corn vary but often are selected by humans for maximum efficiency and productivity of the corn plant in its expected environment. The host corn plant is selected for many characters related to grain productivity, including genetic uniformity such as with single cross hybrids. Corn genetics usually includes production pathogen defense systems that are reactive to avoid needless wasteful, unneeded anti- pathogen compounds. Genetics for resistance requires recognition of a pathogen invasion plus genetics to produce the anti-pathogen chemicals. We are fortunate that corn diversity during its 10,000-year history has allowed existence of genetics for resistance to all potential pathogens somewhere in the corn genome. But within any farmer’s single cross field usually there is only one set of genes and if the hybrid is especially productive it may be common in many fields for a 5-10-year span. Below is my summary of the known interactions with most of the new diseases that have occurred in that last 40+ years in the USA. It is notable that wide use of corn on corn (COC), minimal tillage and susceptible genetics has led to the increase in each of these diseases to the point of being concerning. Resistance was identified, reducing the more severe threat, but some environmental aspects are difficult to change because of other beneficial factors. Dynamics of pathogen genetics, environment and corn genetics are in constant change, requiring us to be constantly alert to emergence of new corn diseases. The pathogen, Cercospora zeae-maydis, had been known for 50 years before it became a major corn disease pest. Two major factors in its emergence as important were increased susceptibility and tillage practices that allowed more diseased leaves from the previous season to remain above ground. Both factors emphasize important dynamics in emergence of new diseases.
An important hybrid parent inbred that became widely used either directly or as a breeding parent in the 1970s was B73. It is susceptible to this pathogen but at the time of its early use this disease was scarce in the U.S.A. corn belt. Concurrent with the introduction of this germplasm, minimum tillage, continuous corn practices were increasing. Both factors were, and still are, important to corn grain production. This fungus cannot compete with soil microorganisms if buried but can overwinter on exposed diseased leaves. When moist, the lesions from the previous season will produces spores (conidia) that are easily picked up by the wind. After landing on a moist corn leaf the spore germinates with hyphae attaching to the epidermal wax surface. It does require a unique accumulation of nearly 100 hours of 90-100% humidity before it will form an appresorium to enzymatically drill into the leaf tissue. This may take only a few days or many days. Initially the disease was prominent in the valleys in humid Eastern U.S.A. but eventually was seen in Ohio River valley. Later it became a major problem in much of the Midwestern USA. Reaching the high humidity is most commonly associated with dew formation, that is not uncommon with evening cooling in the Midwest and especially in corn fields. The resistance factor was also an important variable. This disease increased with susceptible hybrids, a few of which were very popular in the late 70’s. Even with low intensity of the spores in tilled fields these few hybrids would show the disease. Most current hybrids will have some lesions in many fields but only occasionally enough to cause significant yield damage. Most ‘new’ corn diseases have a similar history. The pathogen is mostly obscure and regarded as insignificant until susceptible genetics becomes widespread and environment changes in some way that favors the pest. With realization that the disease is significant, corn breeders tap the vast genetic diversity of maize to identify and incorporate genes for moderation of the pathogens effect on corn grain production. Often those genetics are available within their breeding program. Genetic diversity among pathogens changes in corn growing environments will forever allow new disease pressures on corn. Thanks to its unique reproductive structure encouraging cross-pollination and its unique history of interactions with humans, resistance to ‘new’ diseases will always be found. All corn pathogens and their host have interacting biology. The rust fungi take the interactions to a higher level. Being obligate (biotrophic) parasites, these fungi can only obtain nutrition through living cells, requiring rust fungi to keep the corn cells alive long enough to absorb photosynthates for reproduction and spreading to more living cells in the same host species and then to finally infect an alternative species to complete its sexual cycle.
The common rust species, Puccinia sorghi, spreads to the most temperate zone corn areas from areas with year around corn, such as tropical area. The means of distribution are urediniospores produced on infected, living corn plants. Easily lifted to high altitude winds, especially with storms moving northward in the USA the spores are deposited in the whorls of young corn plants. Requiring only 6 hours of moisture, the germinating spores infect through the stomata of newly emerging leaves. From there hyphae produce an haustoriium that penetrates the cell wall but not the cell membrane, allowing the corn cell to continue with photosynthesis. Instead of the resulting photosynthates moving elsewhere in the plant, it is absorbed by the rust fungus. The time from infection to pustule formation varies with the host genetics but it is only a matter of days before the fungus produces the red urediniospores and spread to other areas on the same plant and elsewhere in the area. Multiple infections of a corn plant, can draw sufficient photosynthates to the fungus to cause light grain weight and total grain loss. Urediniospores of Puccinia sorghi only infect corn. As the pustule ages, perhaps because the host cells die, the fungus produces black spores (teliospores) that can only infect Oxalis (shamrock) species of plants. After infecting the Oxalis plant, the fungus undergoes sexual reproduction and eventually produces another spore type (aeciospores) that can infect corn. However, with Oxalis plants not living through the winter in most temperate area, this part of the fungus life cycle is only completed in tropic and semi-tropic regions. It is there that maximum genetic variability in the fungus occurs. Also in those areas, the cycle is completed with production of new urediniospores that are carried by winds to temperate areas. Consequently, corn infection in temperate areas is dependent upon movement of urediniospores from where corn is grown most of the year. Resistance by the corn plant must either limit the successful penetration through the stomata and/or successful penetration of the cells. This results in few pustules on the leaves. This general or horizontal resistance is controlled by many genes and is considered stable against many races of this pathogen. A more complete resistance resulting in no pustules starves the fungus by causing the host cell to collapse and therefore produce no nutrition for the rust fungus. No pustules develop if this gene is effective against the attack race of the fungus. This vertical resistance is controlled by a single gene. But, of course, as is common with single gene resistance systems, a race of the pathogen can overcome the resistance. Several single genes in corn have been identified and likewise several races of Puccinia sorghi have been identified that can overcome each of the resistance genes. Natural selection for preferable genetics works for pathogens and their hosts! Resistance to leaf diseases in corn that limits the number and size of lesions is called horizontal or quantitative resistance and usually involves 3-5 genes. Ratings for this type of resistance involve a scale developed after considering disease pressure from different environments but does provide a stable type of resistance. Resistance to some leaf pathogens can have another system, usually involving a single gene, that stops some races of a pathogen more completely than horizontal resistance. This latter type of resistance is called vertical or qualitative resistance.
The Ht1 gene was discovered in 1964. It prevented the northern leaf blight pathogen, Exserohilum turcicum, from developing the normal wilted lesion symptom after it reached the vascular tissue. Instead of the normal lesion formation, a small yellow streak developed and, most importantly, the fungus failed to reproduce with spores capable of spreading the disease within the corn field. This seemed ideal in the USA because its presence was easily identified and damage from the disease was eliminated. Consequently, the Ht1 gene was utilized in commercial hybrids during the 1970’s. In 1979, a race of this fungus was found in several locations in the U.S. Corn Belt that overcame the Ht1 gene resistance, resulting in normal lesion and sporulation. It appears that the fungal gene responsible to overcome the Ht1 gene was present in a low frequency within the widespread population of E. turcicum. Its frequency increased as it gained competitive advantage over those individuals without this gene. Similar races of this fungus had already been noted in South Africa and the Philippines. Other single genes ( Ht2, Ht3, Htn) for resistance to E. turcicum have also been identified, as well as races of the fungus that overcome those resistance systems. This is not a new phenomenon. Genetic diversity within pathogens have repeatedly shown an increase of individual genes producing products to overcome single gene resistance. It should be noted that the term race for a pathogen refers to only a single gene difference within the pathogen population. It probably existed as a mutation, allowing a slight structural change in a protein that happened to be attacked by the host plant’s resistance product. Qualitative, vertical resistance to a disease in corn offers quick answers but stable, long-term benefits are best when quantitative, horizontal involving several genes are employed in corn hybrids. Leaf epidermal cells walls and the waxy leaf surface provide the first line of defense against microbes. Pathogens adapted to overcoming this defense set off the next defense system after penetrating the leaf. This is initiated by the plant detecting the presence of the intruder. Plant cells nearby detect the presence of a protein exuded by the pathogen. Such proteins are called effectors, as they are detected chemically by host cells near the invader. Upon detection, these adjacent host cells produce potential microbe-inhibiting compounds such as reactive oxygen, nitric oxide, specific enzymes, salicylic acid and other hormones to effectively thwart the pathogen growth. Much initial reaction is limited to host cells adjacent to the infection site.
Resistance to corn leaf pathogens such as Exserohilum turcicum, cause of northern leaf blight, Cercospora zeae-maydis (gray leaf spot) and Bipolaris maydis (southern corn leaf blight) Involve detection of that specific pathogen and production of more general antimicrobial products in the immediate area of the pathogen. These two steps are inherited independently. Perhaps the pathogen detection system is more specific to the pathogen, accounting for a corn variety being more resistant to one pathogen than another. On the other hand, I am suspicious that if two pathogens arrive in the same area of the plant, only one will survive, as if the plant reacts to the first one by producing general resistance compound that inhibit the infection by the second one to arrive in the same area. The system described above is referred to as general or horizontal resistance. It is controlled by 3-5 genes for products to detect and reduce spread of the pathogen. Horizontal resistance is expressed in corn plants by fewer leaf disease lesions. Evaluation of varieties for this type of lesion has some ambiguity however, because the number of lesions or amount of leaf damage is also affected by the intensity of disease pressure. Heavily diseased leaves from the previous season in fields of low tillage, with frequent early season rain can result in more leaf lesions in a variety of good general resistance to a pathogen than will occur in one of poor resistance with little disease pressure. Characterization of horizontal resistance level to a pathogen requires a rating scale that has some consideration of disease pressure and relativity to other varieties. It is best done when each variety is exposed to the same pathogen intensity at the same stage of leaf maturity. Differences expressed as lesion numbers, size of lesions and percent of leaf destruction can be used to indicate the level of general resistance to that pathogen. I prefer to make ratings based upon several plants exposed to the pathogen in what I project to be somewhat heavy disease pressure in most USA corn environments. With artificial exposure to the pathogen by placing spores in the plant whorl, each plant receives more-or-less the same pressure. Expression of resistance will show 1-2 weeks later. Those varieties with abundance of larger lesions are deemed more susceptible than those with fewer and often smaller lesions. Consequently, it is assumed that will simulate the reactions in fields with somewhat heavy pressure from that pathogen. Any evaluation of horizontal resistance includes consideration of disease pressure and relativity to other varieties. Exserohilum turcicum remains alive, but dormant, in diseased, dead leaf tissue on the soil. When temperatures reach 60-80°F, and the leaf is wet for 24-48 hours, this fungus produces hundreds of spores (conidia) on the surface of the old northern leaf blight lesion. Dispersed by wind, these conidia that land on wet corn leaf surfaces will germinate with hyphae, forming appresoria, attach to the leaf surface, send out a penetration peg and enter the leaf epidermal cells within a few hours. The leaf whorl of a young corn plant is a common site for this infection because of the moisture.
The plant tissue near the site of infection reacts to the presence of the intruder by producing anti-fungal products. These biochemicals not only inhibit growth of this fungus but cause a small chlorotic spot (2-3 mm in diameter) within 48 hours of the infection. This may be sufficient to stop the fungus from progressing in the leaf but with infection by large numbers of spores, a few hyphae can progress on towards the vascular tissue of the leaf. Once inside the xylem the hyphae essentially fill the tissue with hyphae, causing a mini-wilt in that immediate area. The typical disease symptom of this disease usually starts to be visible about 10 days after infection, as a grey, elongate lesion. These lesions mature about a week later, the fungus eventually stopped from further growth in the corn plant but, with moisture, stimulated to produce more conidia within the lesion anytime during or after the season. Leaf vascular systems, with multiple branches, are not significantly harmed by a few of these disease lesions but large numbers of lesions will take its toll on movement of sugars in the phloem from the leaves to the roots and developing kernels. Corn varieties vary in the ability to stop Exserohilum turcicum from reaching the vascular tissue of infected leaves. This resistance system, controlled by 3-4 genes, affects the number of lesions that develop. There are also single, dominant genes that cause specific reactions to the fungus. These genes are designated as Ht1, Ht2, Ht3 and HtN. The first three of these interact after the fungus has reached the vascular tissue, causing a chlorotic streak instead of the gray wilted lesion and inhibits most of the fungus’s ability to produce conidia. The Ht1 gene was effective in the USA for most of the 1970’s, but typical of most single gene resistance systems, the population of the fungus included some individuals with a gene to overcome this resistance. Eventually it became evident that there were races of the Exserohilum turcicum that could overcome each of the single genes for resistance. The most stable resistance is the multi-genic one. Dynamics of spore intensity, leaf surface moisture and corn genetic resistance affect whether this disease affects performance of the crop. Heavy damage usually results in early death of root tissue because of lack of photosynthates reaching the tissues. Early death of root tissue causes general plant wilting, shutting off translocation of sugars to the developing kernels. Thus, the biggest damage from this leaf disease come in the form of stalk lodging and light grain weight. Before discussing the fungi causing corn leaf diseases it may be helpful to address the confusing names given to the pathogens. International standards, allowing for research communication across languages, was established in 1753 by Carl Linnaeus. Every known organism is assigned a Latin-based name for genus and species. Species that are determined to be closely related are assigned a common genus name. Furthermore, there is an International Code of Botanical Nomenclature in which there is a traditional agreement on the name.
Most plant species have traditionally been designated names and relationships based upon structures of flowers, with the assumption that this sexual reproduction stage would be genetically stable. This standard has been more difficult to apply to many fungi, including those that attack corn plants. Many of those have two types of reproductions. The asexual reproduction is often dominant as the spores (conidia) are produced without sexual recombination and, in some cases, are the only reproduction form of the fungus that is known. Consequently, a name was created based on the conidia for the corn pathogen causing northern leaf blight as Helminthosporium turcicum in 1876. Because the conidia were long and dark, like a worm (Helminth) it and a similarly shaped fungus causing southern corn leaf blight (Helminthosporium maydis) were assumed to be in the same genus. In 1959, these two fungi were separated from other members of the genus Helminthosporium into a separate genus named Bipolaris. These names were still based upon conidia structure. Later, again based upon conidia structure, the genus was changed to Drechslera in 1966. In 1974, the northern leaf blight pathogen was given the name of Exserohilum turcicum, because of the microscopic structure of the conidia that is like a group of related fungi. Also in 1974, the rare sexual stage of this species was identified as being like others in the genus Setosphaeria. The sexual name generally takes precedence and thus the ‘official’ name should be Setosphaeria turcica. Formal literature concerning the cause of northern leaf blight will list the most recently accepted name first as well as the authors of the research giving it the name. The formal name for this fungus is Setosphaeria turcica (Luttr.) K.J. Leonard& Suggs. (syns. Bipolaris turcica (Pass.) Shoemaker. Drechslera turcica (Pass.) Subram. &P. C. Jain. Helminthosporium turcicum Pass. Trichometasphaereia turcica Luttr.) These name histories, and the taxonomy features involved are of interest to those who study fungi per se (I was one) but those who are interested in diseases of corn mostly want the cause to be identified and do not want to be confused by the names. On one hand, we appreciate that the name correctly identifies the genetic relationships with other fungi and that it could lead to further understanding of its control, but on the other hand we do not want to be confused by the name changes. I like Helminthosporium turcicum but do adapt to Exserohilum turcicum in my communications with others in the practical aspects of corn. I cannot escape my mycological past, however, so the Latin names for a species will be italicized. |
About Corn JournalThe purpose of this blog is to share perspectives of the biology of corn, its seed and diseases in a mix of technical and not so technical terms with all who are interested in this major crop. With more technical references to any of the topics easily available on the web with a search of key words, the blog will rarely cite references but will attempt to be accurate. Comments are welcome but will be screened before publishing. Comments and questions directed to the author by emails are encouraged.
Archives
December 2021
Categories
|
 RSS Feed
RSS Feed